
Cynthia Wolberger
WBSB 606
Research Interests
Protein function is dynamically regulated in the cell by reversible posttranslational modifications. We are interested in the mechanism by which chromatin modifications regulate transcription, nucleosome dynamics and the response to DNA damage. A particular focus is on the non-degradative signaling played by the attachment of the small protein, ubiquitin, to histone proteins.
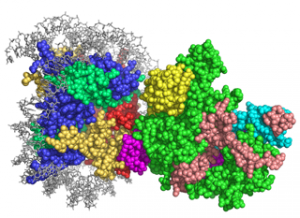
We study the role that ubiquitination plays in these processes and the mechanism of cross-talk between histone ubiquitination, acetylation and methylation, which together orchestrate the complex events underlying mRNA transcription and DNA repair. We use a combination of cryo-electron microscopy, x-ray crystallography, solution biochemistry, cell-based assays, and a variety of biophysical tools to discover the mechanisms underlying these essential cellular processes. The insights from our structural studies are being used to develop novel therapeutic agents that specifically target enzymes that drive a variety of cancers.
Selected Publications
Zhao F, Hicks CW, Wolberger C. (2023) Mechanism of histone H2B monoubiquitination by Bre1. Nature Structural and Molecular Biology, 30(11):1623-1627
Rahman S, Hoffmann NA, Worden EJ, Smith ML, Namitz KEW, Knutson BA, Cosgrove MS, Wolberger C. (2022) Multistate structures of the MLL1-WRAD complex bound to H2B-ubiquitinated nucleosome. Proc Natl Acad Sci U S A. 119(38):e2205691119.
Morgan M, Ikenoue T, Suga H, Wolberger C. (2022) Potent macrocycle inhibitors of the human SAGA deubiquitinating module. Cell Chemical Biology 29(4):544-554.e4.
Worden EJ, Zhang X, Wolberger C (2020) Structural basis for COMPASS recognition of an H2B-ubiquitinated nucleosome. Elife 9. pii: e53199. doi: 10.7554/eLife.53199.
Worden EJ, Hoffmann N, Wolberger C. (2019) Mechanism of cross-talk between H2B ubiquitination and H3 methylation by Dot1L. Cell, 176(6):1490-1501.
Nune M, Morgan MT, Connell Z, McCullough L, Jbara M, Sun H, Brik A, Formosa T, Wolberger C. (2019) FACT and Ubp10 collaborate to modulate H2B deubiquitination and nucleosome dynamics. eLife, pii: e40988. doi: 10.7554/eLife.40988
Morgan M, Haj-Yahya M, Ringel AE, Bandi P, Brik A, Wolberger C (2016) Structural basis for histone H2B deubiquitination by the SAGA DUB module. Science 351(6274): 725-8.
